Edwards S. Boyden Massachusetts Institute of Technology Title: “Ultra-Multiplexed Nanoscale In Situ Proteomics for Understanding Synapse Types” BRAIN category: Tools for Cells and Circuits (RFA MH-14-216)
Dr. Boyden’s team will simultaneously image both the identities and locations of multiple proteins within individual synapses – made possible by a new technique called DNA-PAINT.
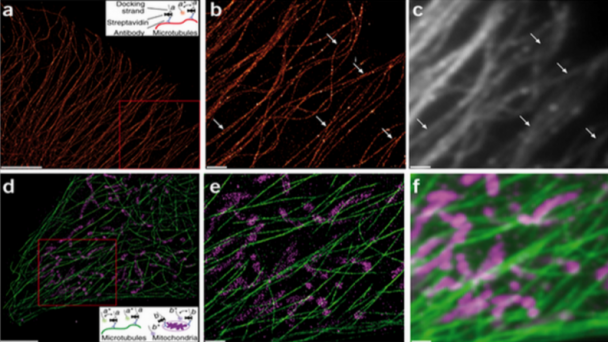
DNA-PAINT super-resolution image of microtubules inside a fixed HeLa cell using Atto 655–labeled imager strands (10,000 frames, 10-Hz frame rate). Inset, labeling and imaging schematic for DNA-PAINT in a cellular environment. From Neuron doi:10.1038/nmeth.2835
Project Description
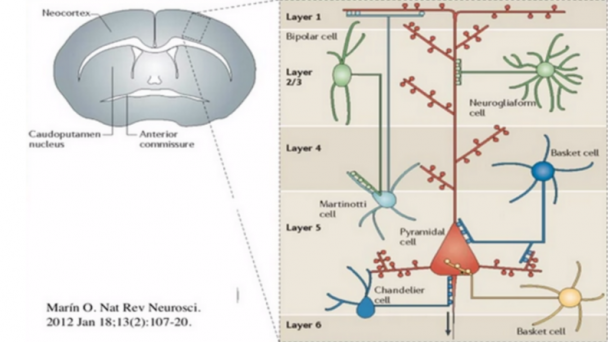
Significant work is ongoing to reveal how different cell types in the brain contribute to behavior and pathology, and how they change in plasticity and disease, empowered by new genetic, optical, and physiological tools. However, the functional activity and dysregulation of neuronal circuits relies critically on the in situ molecular composition of neuronal synapses. Although it is clear that the properties of a given synapse are determined by, amongst other things, the specific types of cells that are thus connected, far less is known about the diversity of synapse types in the brain than cell types, perhaps because this is an intrinsically proteomic problem: a given neuron might make many ...
OnAir Post: Multiplexed Nanoscale In Situ Proteomics