Associate Professor of Biology
Director, Kanold Lab
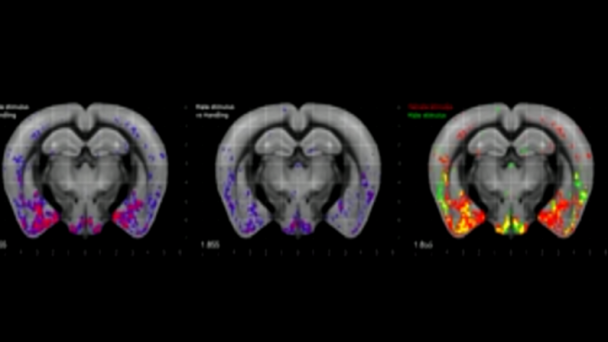
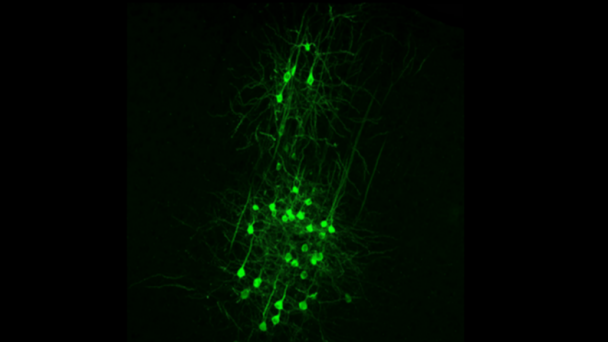
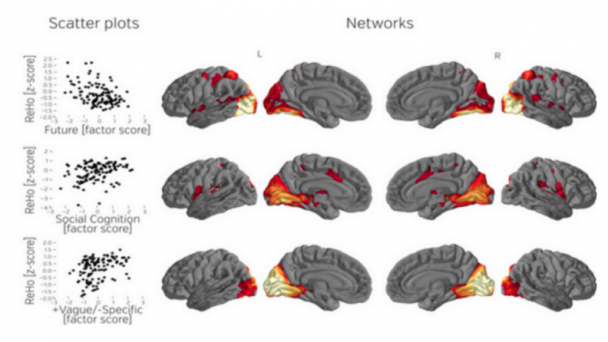
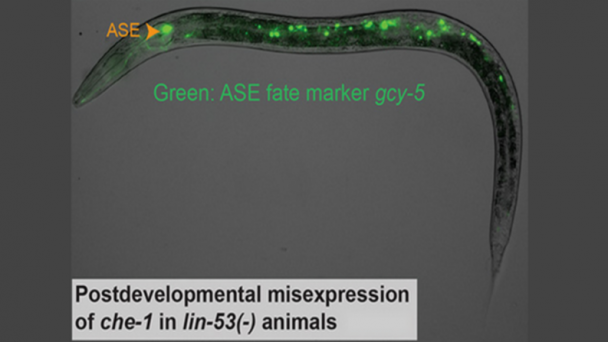
Dr. Kanold studies the development and plasticity of the brain, in particular how periods of learning and plasticity are initiated and controlled. His work focuses on the development of the central auditory and visual system in particular on the role of early cortical circuits in brain wiring. He uses advanced neurophysiological, in vivo imaging, optogenetic, molecular and computational techniques.
Web Information
Webpage: biology.umd.edu/patrick-kanold.html
UMD Neuroscience and Cognitive Science
BRAIN Initiative Grant – “Crowd coding in the brain: 3D imaging and control of collective neuronal dynamics”
Contact Information
Email: pkanold@umd.edu
Phone: 301.405.5741
Address: 1116 Bioscience Research Building
College Park, MD 2074
Biography
Awards
2007 Ralph E Powe Award
2010 Alfred P. Sloan Foundation
2013 NOHR/ARo Burt Evans Award
Education
Dipl. Ing (M.Sc.), Technische Universität Berlin, Germany, 1994
Ph.D., Johns Hopkins University, 2000
PostDoc, Harvard Medical School 2000-2005
Instructor, Harvard Medical School 2005-2006
Research
Dr. Kanold studies the development and plasticity of the brain, in particular how periods of learning and plasticity are initiated and controlled. His work focuses on the development of the central auditory and visual system in particular on the role of early cortical circuits in brain wiring. He uses advanced neurophysiological, in vivo imaging, optogenetic, molecular and computational techniques. His work furthers our understanding of how prenatal and postnatal brain injury ...